Simulation Examples#
import equinox as eqx
import jax.numpy as jnp
import jax.random as rng
import matplotlib.pyplot as plt
import stochastix as stx
from stochastix.utils.visualization import plot_abundance_dynamic
key = rng.PRNGKey(42)
# jax.config.update('jax_enable_x64', True)
plt.rcParams['font.size'] = 18
/Users/francesco/Documents/GitHub/stochastix/stochastix/utils/__init__.py:3: FutureWarning: The 'stochastix.utils.optimization' module is experimental and it may change or be removed without notice in future versions.
from . import nn, optimization, visualization
Oscillator#
# Define the oscillator reactions using the new API
reactions = [
stx.Reaction('X0 + X1 -> 2 X1', stx.kinetics.MassAction(0.01)),
stx.Reaction('X1 + X2 -> 2 X2', stx.kinetics.MassAction(0.01)),
stx.Reaction('X0 + X2 -> 2 X0', stx.kinetics.MassAction(0.01)),
stx.Reaction('2 X0 -> X0', stx.kinetics.MassAction(0.0001)),
stx.Reaction('2 X1 -> X1', stx.kinetics.MassAction(0.0001)),
stx.Reaction('2 X2 -> X2', stx.kinetics.MassAction(0.0001)),
]
model = stx.ReactionNetwork(reactions)
print(model)
R0: X0 + X1 -> 2 X1 | MassAction
R1: X1 + X2 -> 2 X2 | MassAction
R2: X0 + X2 -> 2 X0 | MassAction
R3: 2 X0 -> X0 | MassAction
R4: 2 X1 -> X1 | MassAction
R5: 2 X2 -> X2 | MassAction
key, subkey = rng.split(key)
x0 = jnp.array([100, 50, 70])
T = 100
sim_results = stx.stochsimsolve(
subkey,
model,
x0,
T=T,
max_steps=int(5e5),
)
print('Time overflow:\t', sim_results.time_overflow)
Time overflow: False
plot_abundance_dynamic(sim_results, time_unit='s');
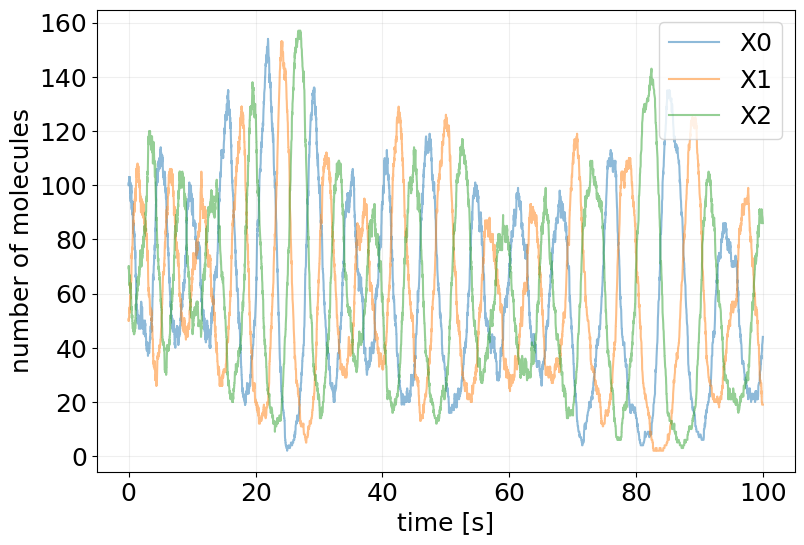
Lotka-Volterra Model (Predator-Prey)#
# Use the built-in Lotka-Volterra generator
lv_model = stx.generators.lotka_volterra_model(
alpha=jnp.array(0.1), # prey reproduction rate
beta=0.001, # predation rate
gamma=0.1, # predator death rate
)
print(lv_model)
R0 (prey_reproduction): prey -> 2 prey | MassAction
R1 (predation): predator + prey -> 2 predator | MassAction
R2 (predator_death): predator -> 0 | MassAction
key, subkey = rng.split(key)
x0 = dict(predator=100, prey=50)
sim_results = stx.stochsimsolve(
subkey,
lv_model,
x0,
T=500,
max_steps=int(5e4),
)
print('Time overflow:\t', sim_results.time_overflow)
Time overflow: False
plot_abundance_dynamic(sim_results, time_unit='s');
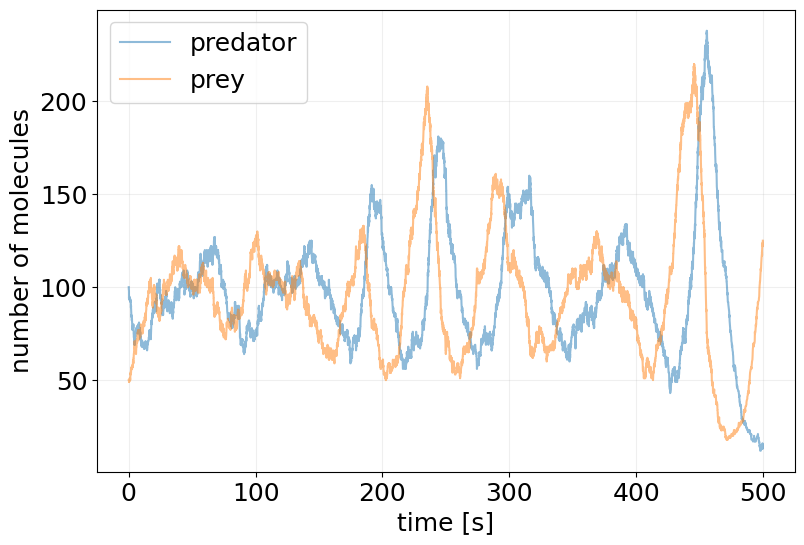
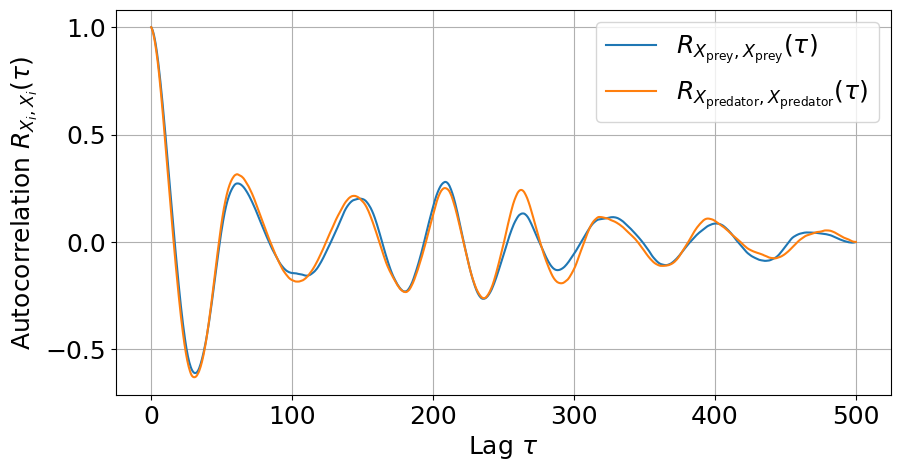
lags, autocorr1 = stx.analysis.autocorrelation(
sim_results, species='prey', n_points=1000
)
lags, autocorr2 = stx.analysis.autocorrelation(
sim_results, species='predator', n_points=1000
)
# Plot autocorrelation
_, ax = plt.subplots(figsize=(10, 5))
ax.plot(lags, autocorr1)
ax.plot(lags, autocorr2)
ax.set_xlabel('Lag $\\tau$')
ax.set_ylabel('Autocorrelation $R_{X_i,X_i}(\\tau)$')
ax.legend(
[
'$R_{X_{\\mathrm{prey}},X_{\\mathrm{prey}}}(\\tau)$',
'$R_{X_{\\mathrm{predator}},X_{\\mathrm{predator}}}(\\tau)$',
]
)
ax.grid(True)
plt.show()
lags, cross_corr1 = stx.analysis.cross_correlation(
sim_results, species1='prey', species2='predator', n_points=1000
)
lags, cross_corr2 = stx.analysis.cross_correlation(
sim_results, species1='predator', species2='prey', n_points=1000
)
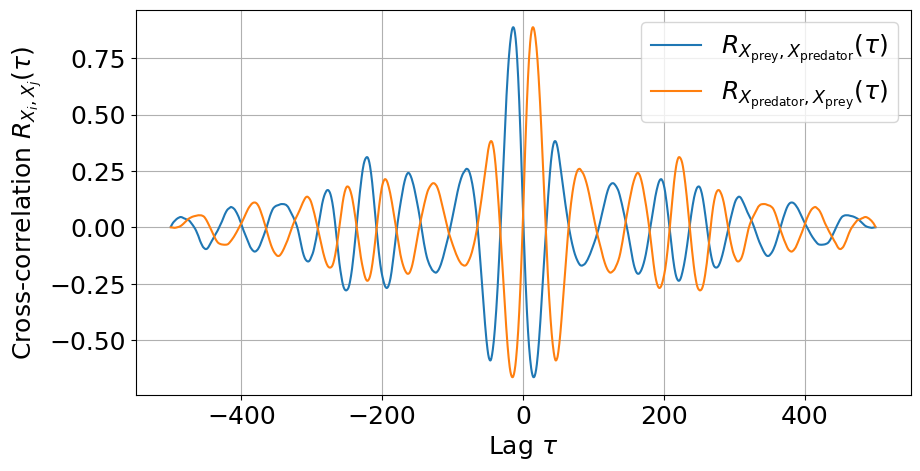
# Plot cross-correlation
_, ax = plt.subplots(figsize=(10, 5))
ax.plot(lags, cross_corr1)
ax.plot(lags, cross_corr2)
ax.set_xlabel('Lag $\\tau$')
ax.set_ylabel('Cross-correlation $R_{X_i,X_j}(\\tau)$')
ax.legend(
[
'$R_{X_{\\mathrm{prey}},X_{\\mathrm{predator}}}(\\tau)$',
'$R_{X_{\\mathrm{predator}},X_{\\mathrm{prey}}}(\\tau)$',
]
)
ax.grid(True)
plt.show()
Test differentiability:
@eqx.filter_jit
def test_loss(network, x0, key):
sim_results = stx.stochsimsolve(
key,
network,
x0,
T=500,
max_steps=int(5e4),
solver=stx.solvers.DifferentiableDirect(),
)
_, cross_corr = stx.analysis.cross_correlation(
sim_results,
species1='prey',
species2='predator',
n_points=1000,
)
return jnp.mean(cross_corr**2)
key, subkey = rng.split(subkey)
eqx.filter_grad(test_loss)(lv_model, x0, key).prey_reproduction.kinetics.k
Array(0.67341566, dtype=float32, weak_type=True)
SIRS Model#
# Use the built-in SIRS generator (with no recovery -> susceptible to get SIR)
sir_model = stx.generators.sirs_model(
beta=0.005, # transmission rate
gamma=0.02, # recovery rate
nu=0.01, # loss of immunity rate (R -> S)
)
print(sir_model)
R0 (infection): I + S -> 2 I | MassAction
R1 (recovery): I -> R | MassAction
R2 (loss_of_immunity): R -> S | MassAction
key, subkey = rng.split(key)
# I, R, S (stored in alphabetical order)
x0 = jnp.array([1, 0, 100])
sim_results = stx.stochsimsolve(
subkey,
sir_model,
x0,
T=300,
max_steps=int(5e5),
)
print('Time overflow:\t', sim_results.time_overflow)
Time overflow: False
sir_model.species
('I', 'R', 'S')
plot_abundance_dynamic(sim_results, time_unit='s', log_x_scale=True);
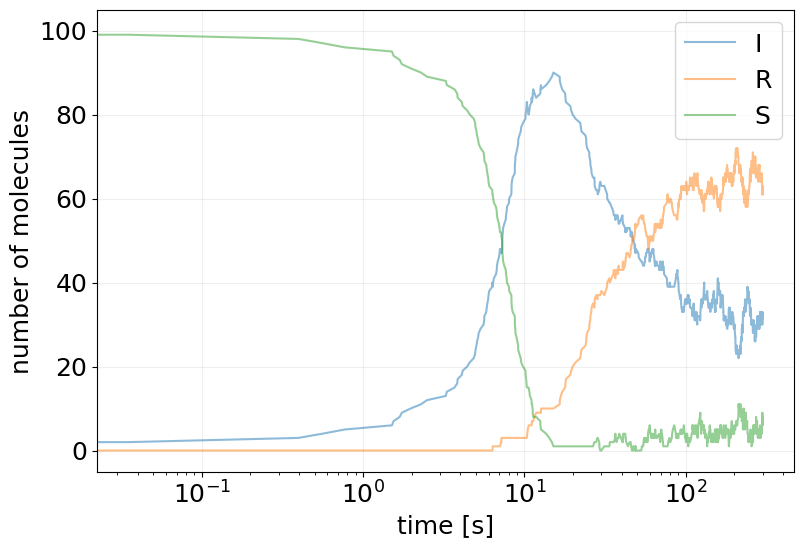
Repressilator#
# Use the built-in repressilator generator
repressilator = stx.generators.repressilator_model(
alpha=30.0, # maximum expression rate
alpha0=0.0, # leaky expression rate
K=30.0, # half-saturation constant
beta=0.1, # degradation rate
n=3.0, # Hill coefficient (cooperativity)
)
print(repressilator)
R0 (gene_A_expression): 0 -> A | HillRepressor
R1 (gene_B_expression): 0 -> B | HillRepressor
R2 (gene_C_expression): 0 -> C | HillRepressor
R3 (protein_A_degradation): A -> 0 | MassAction
R4 (protein_B_degradation): B -> 0 | MassAction
R5 (protein_C_degradation): C -> 0 | MassAction
key, subkey = rng.split(key)
x0 = jnp.ones(repressilator.n_species) * 10
sim_results = stx.stochsimsolve(subkey, repressilator, x0, T=300.0, max_steps=int(5e5))
print('Time overflow:\t', sim_results.time_overflow)
Time overflow: False
plot_abundance_dynamic(sim_results, time_unit='s');
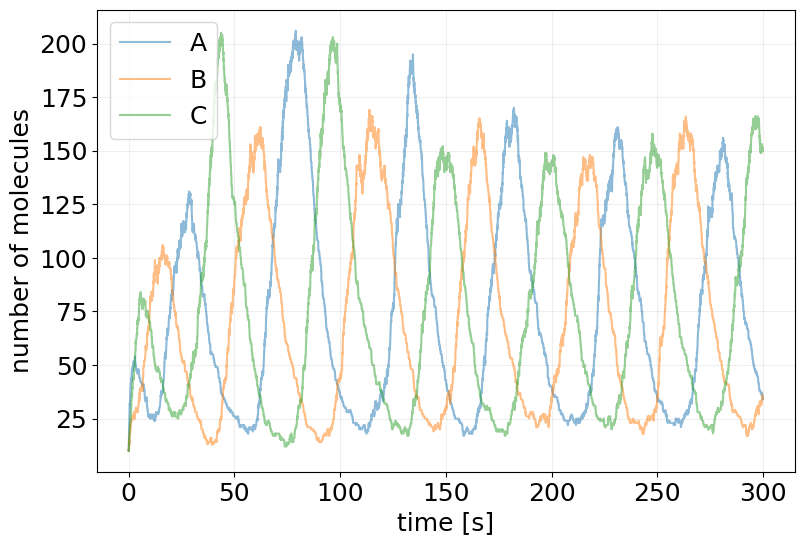
C2-FFL - Coherent Type 2 Feed-Forward Loop#
from stochastix import Reaction, ReactionNetwork
from stochastix.kinetics import HillActivator, HillRR, MassAction
c2_ffl_reactions = [
Reaction(
'0 -> Y',
HillActivator(regulator='X', v=50.0, K=20.0, n=3.0),
name='Y_production',
),
Reaction(
'0 -> Z',
HillRR('X', 'Y', v=50.0, K1=40.0, K2=5.0, n1=3.0, n2=3.0, logic='and'),
name='Z_production',
),
Reaction('Y -> 0', MassAction(k=1.0), name='Y_degradation'),
Reaction('Z -> 0', MassAction(k=1.0), name='Z_degradation'),
]
c2_ffl = ReactionNetwork(c2_ffl_reactions)
print(c2_ffl)
R0 (Y_production): 0 -> Y | HillActivator
R1 (Z_production): 0 -> Z | HillRR
R2 (Y_degradation): Y -> 0 | MassAction
R3 (Z_degradation): Z -> 0 | MassAction
Static Inputs#
key, subkey_high, subkey_low = rng.split(key, 3)
# x0_high = jnp.array([50.0, 0.0, 0.0])
x0_high = dict(X=50.0, Y=0.0, Z=0.0)
# x0_low = jnp.array([5.0, 0.0, 0.0])
x0_low = dict(X=5.0, Y=0.0, Z=0.0)
T = 100.0
sim_high = stx.stochsimsolve(subkey_high, c2_ffl, x0_high, T=T, max_steps=int(5e5))
sim_low = stx.stochsimsolve(subkey_low, c2_ffl, x0_low, T=T, max_steps=int(5e5))
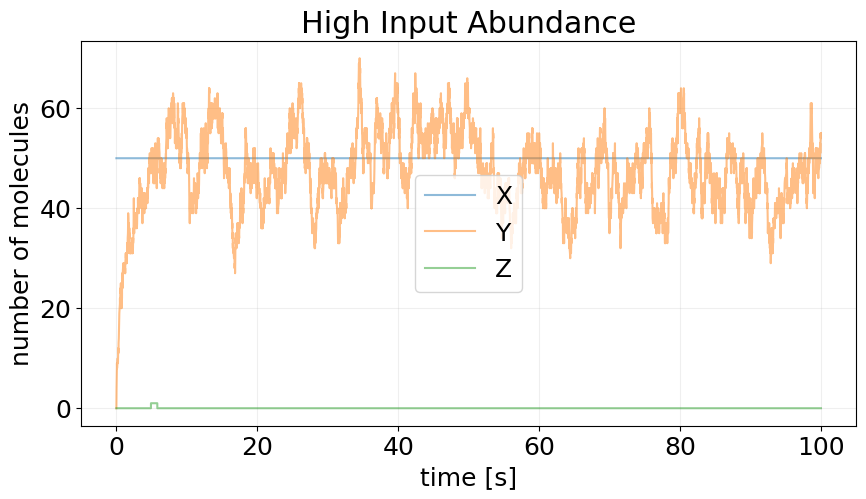
_, ax = plt.subplots(figsize=(10, 5))
plot_abundance_dynamic(sim_high, time_unit='s', ax=ax)
ax.set_title('High Input Abundance')
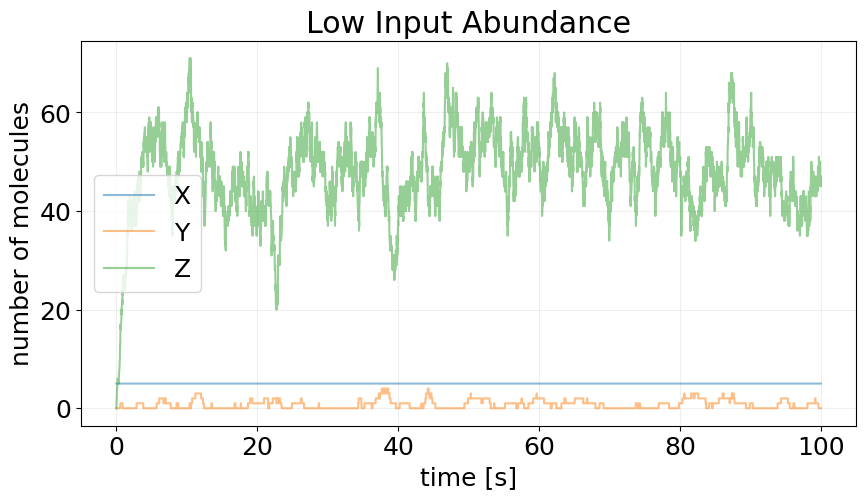
_, ax = plt.subplots(figsize=(10, 5))
plot_abundance_dynamic(sim_low, time_unit='s', ax=ax)
ax.set_title('Low Input Abundance');
Dynamic Input (Time-controlled species)#
from stochastix.controllers import Timer
controlled_species = 'X'
time_triggers = jnp.array([20.0, 60.0, 80.0])
species_t = jnp.array([0.0, 50.0, 15.0])[:, None]
timed_controller = Timer(controlled_species, time_triggers, species_t)
timed_controller
Timer(
controlled_species=('X',),
time_triggers=(20.0, 60.0, 80.0),
species_at_triggers=((0.0,), (50.0,), (15.0,))
)
key, subkey = rng.split(key)
x0_high = jnp.array([50.0, 0.0, 0.0])
sim_ctrl = stx.stochsimsolve(
subkey_high,
c2_ffl,
x0_high,
T=100.0,
controller=timed_controller,
max_steps=int(5e5),
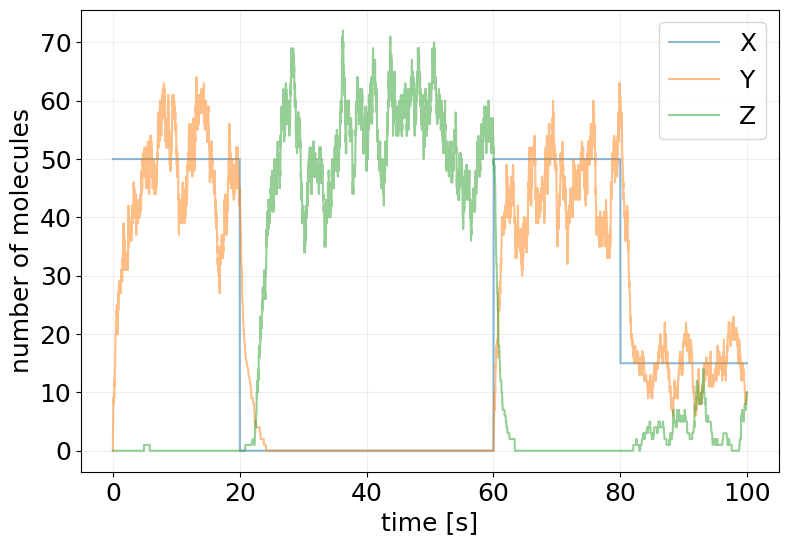
)
plot_abundance_dynamic(sim_ctrl, time_unit='s');