import jax
import jax.numpy as jnp
import jax.random as rng
import matplotlib.pyplot as plt
import stochastix as stx
from stochastix.utils.visualization import plot_abundance_dynamic
jax.config.update('jax_enable_x64', True)
key = rng.PRNGKey(10)
plt.rcParams['font.size'] = 18
/Users/francesco/Documents/GitHub/stochastix/stochastix/utils/__init__.py:3: FutureWarning: The 'stochastix.utils.optimization' module is experimental and it may change or be removed without notice in future versions.
from . import nn, optimization, visualization
Chemical Oscillator Model
# Define the oscillator reactions using the new API
reactions = [
stx.Reaction('X0 + X1 -> 2 X1', stx.kinetics.MassAction(0.01)),
stx.Reaction('X1 + X2 -> 2 X2', stx.kinetics.MassAction(0.01)),
stx.Reaction('X0 + X2 -> 2 X0', stx.kinetics.MassAction(0.01)),
stx.Reaction('2 X0 -> X0', stx.kinetics.MassAction(0.0001)),
stx.Reaction('2 X1 -> X1', stx.kinetics.MassAction(0.0001)),
stx.Reaction('2 X2 -> X2', stx.kinetics.MassAction(0.0001)),
]
oscillator = stx.ReactionNetwork(reactions)
R0: X0 + X1 -> 2 X1 | MassAction
R1: X1 + X2 -> 2 X2 | MassAction
R2: X0 + X2 -> 2 X0 | MassAction
R3: 2 X0 -> X0 | MassAction
R4: 2 X1 -> X1 | MassAction
R5: 2 X2 -> X2 | MassAction
Stochastic Simulation
key, subkey = rng.split(key)
x0 = jnp.array([100, 50, 70])
T = 100
ssa_results = stx.stochsimsolve(
subkey,
oscillator,
x0,
T=T,
max_steps=int(5e5),
)
print('Time overflow:\t', ssa_results.time_overflow)
plot_abundance_dynamic(ssa_results);
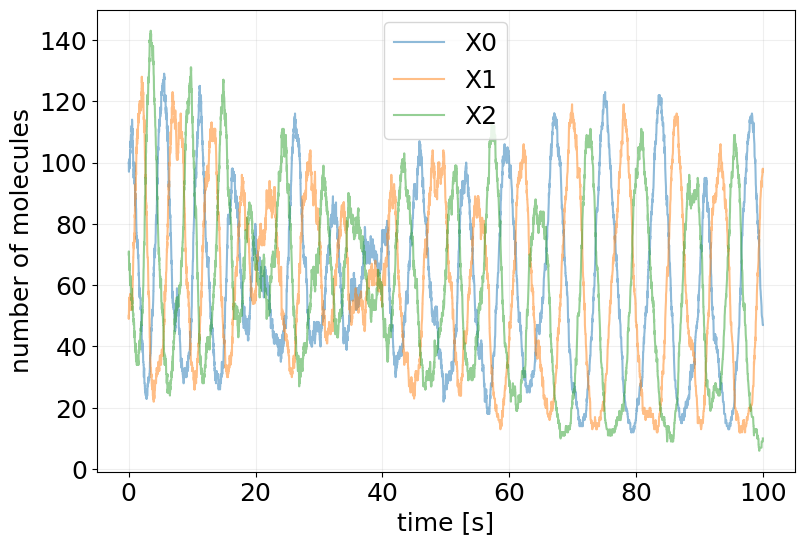
Conversion to ODE Model
from diffrax import Dopri5, ODETerm, SaveAt, diffeqsolve
ode_term = ODETerm(oscillator.vector_field)
# or
# ode_term = oscillator.diffrax_ode_term()
dt0 = 0.1
saveat = SaveAt(ts=jnp.linspace(0.0, T, 1000))
ode_results = diffeqsolve(
ode_term, Dopri5(), t0=0.0, t1=T, dt0=dt0, y0=x0, saveat=saveat, max_steps=int(1e6)
)
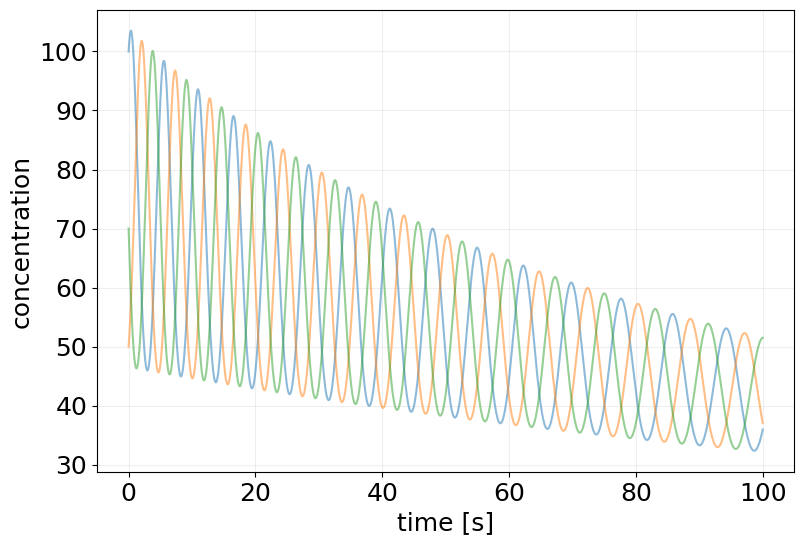
plt.figure(figsize=(9, 6))
plt.plot(ode_results.ts, ode_results.ys, alpha=0.5)
plt.xlabel('time [s]')
plt.ylabel('concentration')
plt.grid(alpha=0.2)
from diffrax import (
ControlTerm,
Euler,
MultiTerm,
ODETerm,
SaveAt,
VirtualBrownianTree,
diffeqsolve,
)
t0, T = 0.0, 100
brownian_motion = VirtualBrownianTree(
t0, T, tol=1e-1, shape=(oscillator.n_reactions,), key=subkey
)
terms = MultiTerm(
ODETerm(oscillator.drift_fn), ControlTerm(oscillator.diffusion_fn, brownian_motion)
)
# or just:
# terms = oscillator.diffrax_sde_term(T, key=subkey)
solver = Euler()
saveat = SaveAt(steps=True)
dt0 = 0.1
sde_results = diffeqsolve(
terms, solver, t0=t0, t1=T, dt0=dt0, y0=x0, saveat=saveat, max_steps=int(1e6)
)
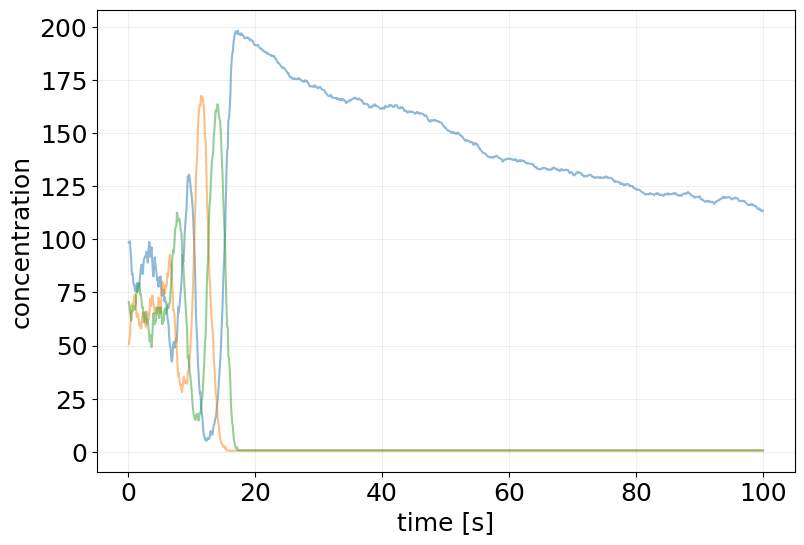
plt.figure(figsize=(9, 6))
plt.plot(sde_results.ts, sde_results.ys, alpha=0.5)
plt.xlabel('time [s]')
plt.ylabel('concentration')
# plt.yscale('log')
plt.grid(alpha=0.2)
Lotka-Volterra Model (Predator-Prey)
# Use the built-in Lotka-Volterra generator
lv_model = stx.generators.lotka_volterra_model(
alpha=0.1, # prey reproduction rate
beta=0.005, # predation rate
gamma=0.05, # predator death rate
)
R0 (prey_reproduction): prey -> 2 prey | MassAction
R1 (predation): predator + prey -> 2 predator | MassAction
R2 (predator_death): predator -> 0 | MassAction
Stochastic Simulation
key, subkey = rng.split(key)
x0 = jnp.array([30, 30])
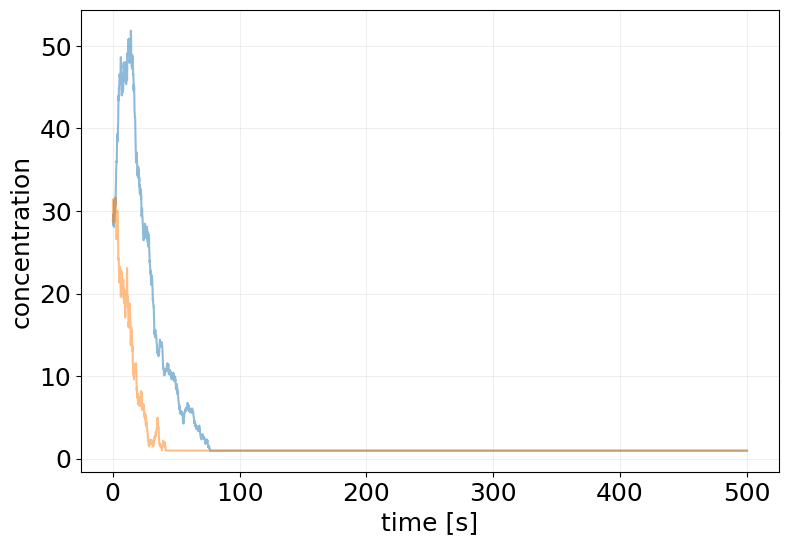
T = 500
ssa_results = stx.stochsimsolve(
subkey,
lv_model,
x0,
T=T,
max_steps=int(5e4),
)
print('Time overflow:\t', ssa_results.time_overflow)
plot_abundance_dynamic(ssa_results, time_unit='s');
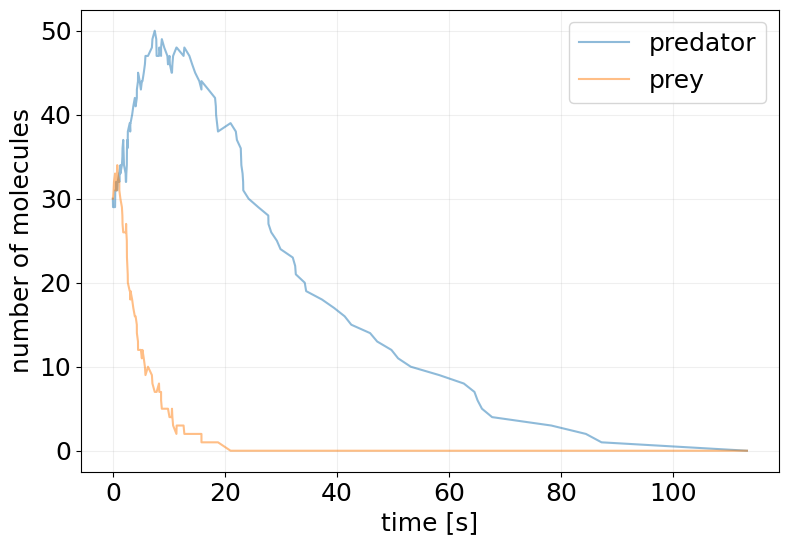
Conversion to ODE Model
from diffrax import Dopri5, ODETerm, SaveAt, diffeqsolve
# ode_term = ODETerm(lv_model.vector_field)
ode_term = lv_model.diffrax_ode_term()
dt0 = 0.1
saveat = SaveAt(ts=jnp.linspace(0.0, T, 1000))
ode_results = diffeqsolve(
ode_term, Dopri5(), t0=0.0, t1=T, dt0=dt0, y0=x0, saveat=saveat, max_steps=int(1e5)
)
plt.figure(figsize=(9, 6))
plt.plot(ode_results.ts, ode_results.ys, alpha=0.5)
plt.xlabel('time [s]')
plt.ylabel('concentration')
plt.grid(alpha=0.2)
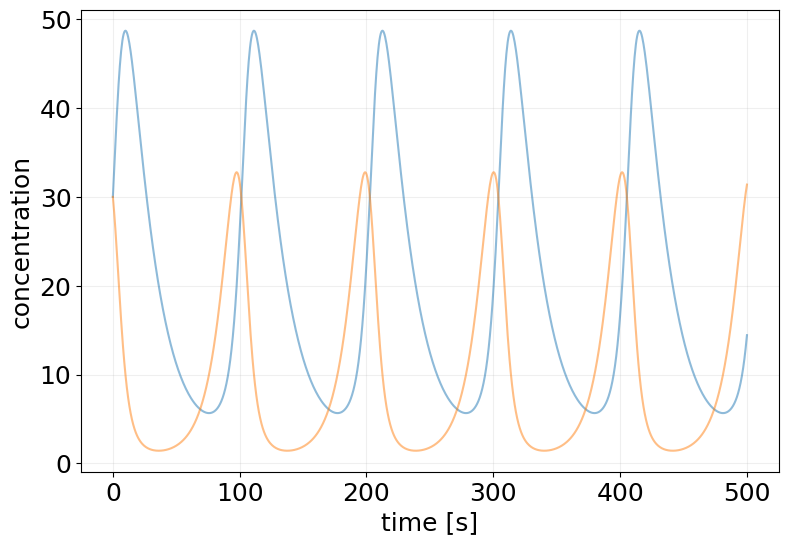
from diffrax import (
ControlTerm,
Euler,
MultiTerm,
ODETerm,
SaveAt,
VirtualBrownianTree,
diffeqsolve,
)
t0, T = 0.0, 500
# brownian_motion = VirtualBrownianTree(
# t0, T, tol=1e-3, shape=(lv_model.n_reactions,), key=subkey
# )
# terms = MultiTerm(
# ODETerm(lv_model.drift_fn), ControlTerm(lv_model.diffusion_fn, brownian_motion)
# )
terms = lv_model.diffrax_sde_term(T, key=subkey)
solver = Euler()
saveat = SaveAt(
steps=True,
)
dt0 = 0.1
sde_results = diffeqsolve(
terms, solver, t0=t0, t1=T, dt0=dt0, y0=x0, saveat=saveat, max_steps=int(1e5)
)
plt.figure(figsize=(9, 6))
plt.plot(sde_results.ts, sde_results.ys, alpha=0.5)
plt.xlabel('time [s]')
plt.ylabel('concentration')
# plt.yscale('log')
plt.grid(alpha=0.2)